Three-dimensional computer reconstruction
3D Reconstruction of Serial
Sections
Dr. Brad Duerstock's research interests involve the development of 3D
surface
reconstructions and volume visualizations of injured segments of spinal
cord from serial histological sections. 3D imaging of spinal cord
injuries was accomplished using different algorithms
that produces either volume visualizations or surface
reconstructions. The software algorithms used for 3D surface
reconstruction were developed at
the Center for Computational
Visualization at The University of Texas.
Isocontouring Method
This surface reconstruction method is similar to Surface Tiling but does
not require circumscribing contours by hand. Object selection is
accomplished by discrimination of pixel values. This method alleviates
labor and time costs and user subjectivity.
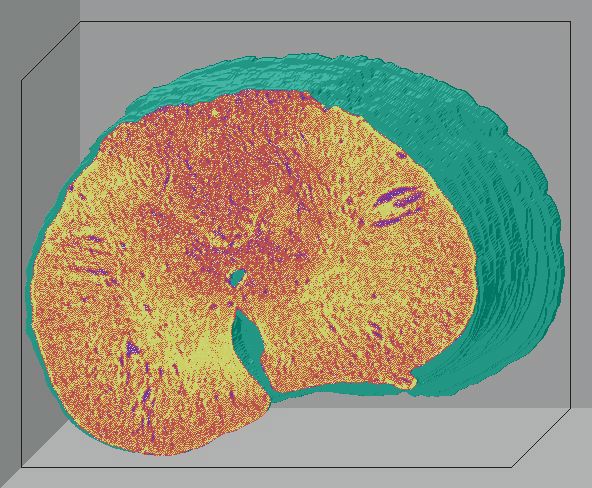
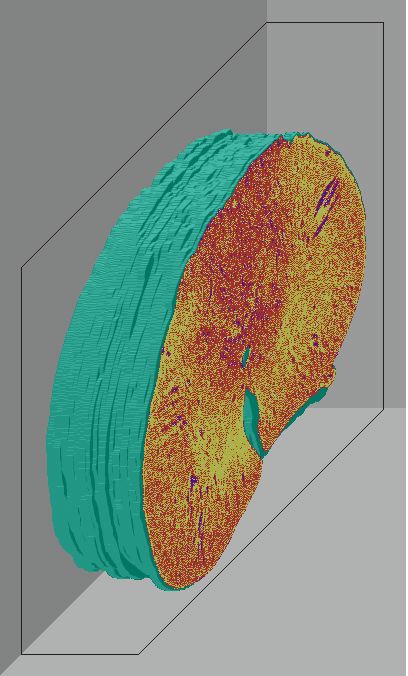
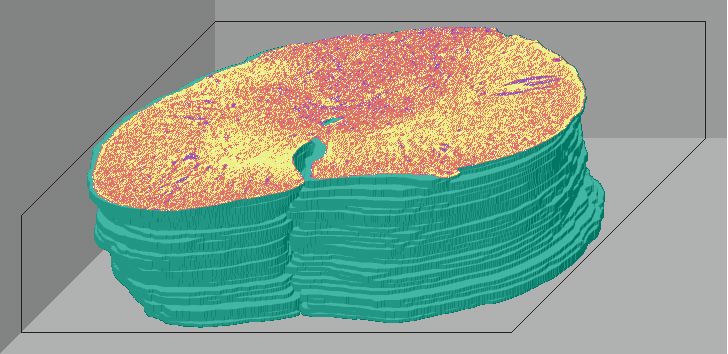
Below are 3D surface reconstructions of a spinal cord segment and its
central grey matter from the lower thoracic level in the rat.
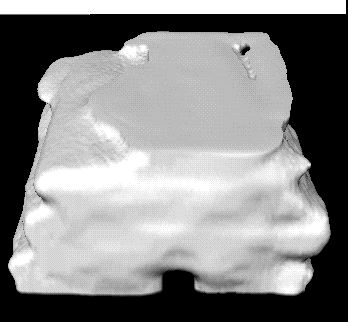
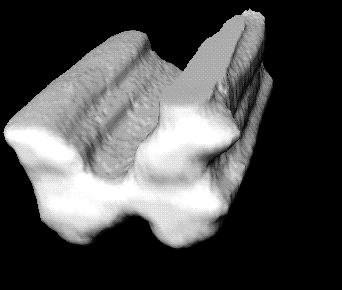 Surface reconstructions not only allow anatomical structures to be viewed
three-dimensionally but also permit surfaces of biological features to
be quantitatively interrogated as well. 3D measurements, such as volume
and surface area, can be precisely calculated for each surface using this
software.
Surface reconstructions not only allow anatomical structures to be viewed
three-dimensionally but also permit surfaces of biological features to
be quantitatively interrogated as well. 3D measurements, such as volume
and surface area, can be precisely calculated for each surface using this
software.
 Animations of the interior of a 6
month-old injured spinal cord segment caused by piercing the cord
during ventral implantation of a piece of muscle. The puncture wound
caused
the formation of 3 large cysts in the chronic injury, unlike the pathology
of compression or contusion injuries.
Animations of the interior of a 6
month-old injured spinal cord segment caused by piercing the cord
during ventral implantation of a piece of muscle. The puncture wound
caused
the formation of 3 large cysts in the chronic injury, unlike the pathology
of compression or contusion injuries.
Walk-through the interior of the SCI (1.6MB .QT)
Navigation through the central canal into one of the
large cysts (700KB .QT)
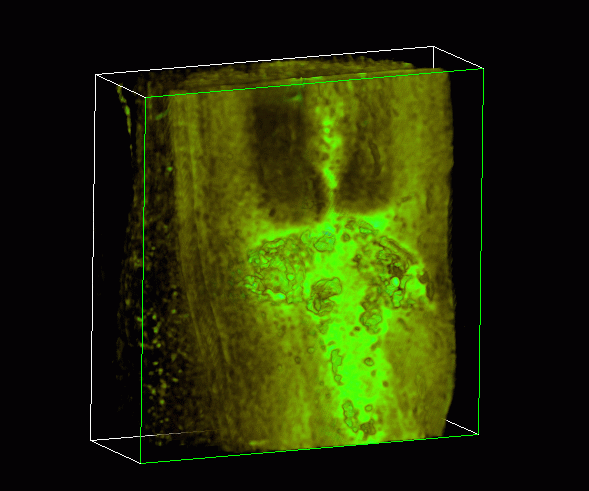
Volumetric Texture Imaging
This is a volume rendering of a three week-old compression SCI. The 4
mm-long segment
of rat spinal cord runs lengthwise from top to bottom. In the center is the
injury site shown in bright green which is characterized by intense
labelling of active macrophages. Macrophages or scavenger cells invade the
subacute injury in large numbers. These cells cause holes or cysts within the
nervous tissue.
 Rotation of the injured spinal cord (751KB .QT)
Rotation of the injured spinal cord (751KB .QT)
Transparency reveals the centralized cysts (1.5MB
.QT)
Virtual cross-sectional view of the spinal segment
reveals the bright green labelling of activated macrophages in the
subacute injury at the center of the cord where damage is greatest.
(940KB .QT)
Surface Tiling Method
This is probably the most common method for 3D surface reconstruction.
This type of algorithm "stacks" 2D contours (manual tracings of regions
of interest) per histological section to form a 3D surface of a particular
anatomical structure.

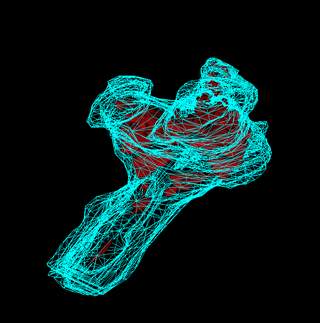
 These 2 images show 3D
surfaces of the injury site (blue) superimposed within histological sections
from which they are generated. This is the same spinal cord as shown by
volumetric texture imaging (above).
These 2 images show 3D
surfaces of the injury site (blue) superimposed within histological sections
from which they are generated. This is the same spinal cord as shown by
volumetric texture imaging (above).
 This method
produces 3D surfaces that are reconstructed
from a series of connected triangles (left). However surface
reconstructions can be textured and made transparent (right) to allow
inspection of internal structures like the grey matter (red). The
ventral side of the uninjured spinal cord segment is facing up.
This method
produces 3D surfaces that are reconstructed
from a series of connected triangles (left). However surface
reconstructions can be textured and made transparent (right) to allow
inspection of internal structures like the grey matter (red). The
ventral side of the uninjured spinal cord segment is facing up.
 The set of histological slices used to compose the
blue 3D reconstructed injury site (937KB .QT)
The set of histological slices used to compose the
blue 3D reconstructed injury site (937KB .QT)
3D lesion (gold) made transparent to reveal the red
cysts within and an island or raft of macrophages (white) floating freely
within one of the largest cysts (1.4MB .QT)
Dynamic navigation within the injury (gold) and
internal cysts (red), perimeter of the spinal cord is shown in blue
(2.6MB .QT)
Publications
Duerstock, B.S. (2003) Double Labeling Serial Sections to Enhance
Three-Dimensional Imaging of Injured Spinal Cord. J Neurosci
Methods 134(1): 101-107.
Duerstock, B.S., Bajaj, C.L., and Borgens, R.B. (2003) A Comparative Study
of the Accuracy of Three-Dimensional Reconstructions of Spinal Cord from
Serial Histological Sections. J Microsc-Oxford 210:138-148 Pt 2.
Duerstock, B.S., Bajaj, C.L., Pascucci, V., Schikore, D., Lin, K.,
Borgens, R.B. (2000) Advances in Three-Dimensional Reconstruction of the
Experimental Spinal Cord Injury. Comput Med Imag Grap 24(6):
389-406.
Moriarty, L.J., Duerstock, B.S., Bajaj, C.L., Lin. K., and Borgens, R.B.
(1998) Two- and Three-Dimensional Computer Graphic Evaluation of the
Subacute Spinal Cord Injury. J Neurol Sci 155: 121-137.
To CPR homepage
Back to Brad's homepage

 Surface reconstructions not only allow anatomical structures to be viewed
three-dimensionally but also permit surfaces of biological features to
be quantitatively interrogated as well. 3D measurements, such as volume
and surface area, can be precisely calculated for each surface using this
software.
Surface reconstructions not only allow anatomical structures to be viewed
three-dimensionally but also permit surfaces of biological features to
be quantitatively interrogated as well. 3D measurements, such as volume
and surface area, can be precisely calculated for each surface using this
software. Animations of the interior of a 6
month-old injured spinal cord segment caused by piercing the cord
during ventral implantation of a piece of muscle. The puncture wound
caused
the formation of 3 large cysts in the chronic injury, unlike the pathology
of compression or contusion injuries.
Animations of the interior of a 6
month-old injured spinal cord segment caused by piercing the cord
during ventral implantation of a piece of muscle. The puncture wound
caused
the formation of 3 large cysts in the chronic injury, unlike the pathology
of compression or contusion injuries.


 These 2 images show 3D
surfaces of the injury site (blue) superimposed within histological sections
from which they are generated. This is the same spinal cord as shown by
volumetric texture imaging (above).
These 2 images show 3D
surfaces of the injury site (blue) superimposed within histological sections
from which they are generated. This is the same spinal cord as shown by
volumetric texture imaging (above).

 This method
produces 3D surfaces that are reconstructed
from a series of connected triangles (left). However surface
reconstructions can be textured and made transparent (right) to allow
inspection of internal structures like the grey matter (red). The
ventral side of the uninjured spinal cord segment is facing up.
This method
produces 3D surfaces that are reconstructed
from a series of connected triangles (left). However surface
reconstructions can be textured and made transparent (right) to allow
inspection of internal structures like the grey matter (red). The
ventral side of the uninjured spinal cord segment is facing up.


